Big Data Challenges
Production portal allows researchers to focus on science
Researchers and clinicians at The Ohio State University’s Wexner Medical Center rely upon the staff members of the Biomedical Informatics Shared Resource (BISR) to address their high-throughput, high-dimensional biological data-analysis needs, such as for next-generation sequencing research. The gigabytes of data generated by these technologies present enormous challenges in storage, computational resources, analysis and interpretation. Moreover, many of the latest technology advances designed to solve big data challenges have not yet been widely adopted by the research and clinical communities.
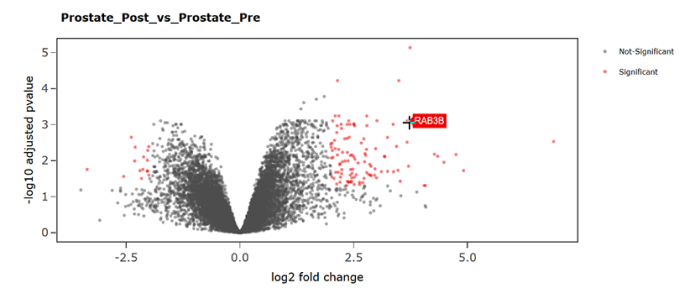
In collaboration with specialists at the Ohio Supercomputer Center (OSC), BISR staff members have developed an interactive analysis reporting system delivered through OSC’s existing OnDemand platform, a secure, scalable, open-source software environment that gives clients web-based access to high performance computing services.
“The cost of supporting this service using OnDemand is lower than building or managing a stand-alone solution,” said Eric Franz, lead engineer for OSC’s Gateways group. “The existing OnDemand installation already has built-in solutions for authentication, authorization, accounting and monitoring that can be leveraged by any OnDemand app.”
“Most importantly, this approach succeeds in shifting the responsibility for ensuring the production availability of the service from BISR to OSC, which enables us to focus more on our domain of work,” said Venkat S. Gadepalli, Ph.D., a research scientist with BISR.
App developers chose to employ the R programming language, considered an ideal tool for data wrangling, analysis and visualization, and Shiny, an R-based library that makes it easy for researchers without the expertise of a software developer to build interactive web apps and reports.
BISR first considered self-hosting Shiny but was concerned about the cost of setting up authentication. The staff also found that authorizing dataset access is a problem that Shiny Server currently does not address. And, because BISR’s results data was already stored on OSC systems, it just made more sense for OSC to host BISR’s Shiny visualization service.
“BISR is actively evaluating and iteratively refining the features of the Shiny report app, which is the main focus for the analyst’s and researcher’s user experience,” explained Maciej Pietrzak, Ph.D., technical director of BISR and a research assistant professor in the biomedical informatics department at Ohio State. “In the next 18 months, we expect to expand the number of users by actively engaging the RNAseq Shiny report app with relevant projects and developing other apps.”
PROJECT LEAD Maciej Pietrzak, Ph.D., The Ohio State University
RESEARCH TITLE Scaling R Shiny Apps to Multiple Concurrent Users in a Secured HPC Environment Using Open OnDemand
FUNDING SOURCE National Science Foundation
WEBSITE medicine.osu.edu/bmi/people/research/maciej_ pietrzak/Pages/index.aspx
FILE UPLOAD: 2019 Research Report_Page17.pdf